
(C) Overlap extension PCR cloning efficiency as a function of the insert length. The number of green colonies was plotted against the number of PCR cycles for each plate. 
coli cells were transformed with 1 μL pQE30/insert overlap extension PCR. (B) Overlap extension PCR cloning efficiency of a gfp gene as a function of the number of PCR cycles. M2, 1 kb DNA ladder M1, assembled plasmid in closed circular and relaxed circular forms. Three nanograms of pQE30 vector were mixed with 175 ng insert (250 molar excess) in 10 μL total volume a 4-μL aliquot of reaction was separated on a 0.8% agarose gel. (A) Products of the overlap extension PCR cloning reaction after 0, 5, 10, 15, 20, 25, and 30 cycles by agarose gel electrophoresis. coli cells.Īnalysis of the overlap extension PCR cloning reaction A small aliquot from each reaction was used to transform E. The original pQE30 vector was destroyed in reactions with DpnI restriction endonuclease ( Figure 1C). Some of the PCR products correspond to the relaxed form of the desirable vector, as revealed by agarose gel analysis ( Figure 2A). High concentrations of the insert and relatively low annealing temperatures in the reaction (5–10☌ below the calculated melting temperature of the primer/plasmid complex) are important for efficient overlap extension. Overlap extension PCR ( Figure 1B) was performed with five different DNA polymerases ( Supplementary Table S1). The gfp gene was PCR-amplified ( Figure 1A) with the chimeric primers (5′ ends complementary to the pQE30 plasmid 3′-end complementary to gfp). We first used gfp for proof-of-principle experiments. 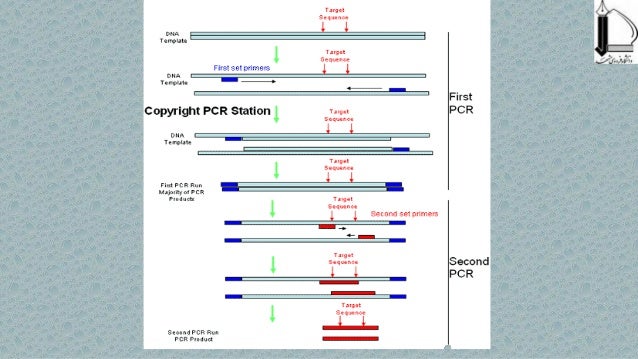
coli after the parental plasmid is destroyed by DpnI digest. (C) The new plasmid can be transformed into E. After several PCR cycles, the new plasmid with two nicks (one on each strand) gets accumulated as a product. (B) Then, vector and insert are mixed, denatured and annealed the hybridized insert then is extended by Phusion DNA polymerase using vector as a template until polymerase reaches 5′ end of the insert. (A) First, the insert is PCR-amplified with the chimeric primers so that the final PCR product has overlapping regions with the vector. We employ the same thermostable polymerase for both PCRs, so inexperienced users can clone efficiently without mastering the idiosyncrasies of multiple restriction enzymes, polymerases, glycosylases, recombinases, and ligases.Īn outline of the overlap extension PCR cloning This relaxed double-stranded plasmid is then transformed into competent Escherichia coli cells, which seal the nicks with DNA repair enzymes ( Figure 1C). Therefore, the final product of the reaction is a double-stranded fusion plasmid with two nicks (one on each strand). F-530 New England BioLabs, Ipswich, MA, USA), crucial for performance of the technique, does not possess strand displacement activity. After denaturation and annealing, the insert strands hybridize to the vector and extend to form new double-stranded plasmid. These extensions subsequently allow the strands of the PCR product ( Figure 1A) to act as a pair of oversized primers on the vector fragment ( Figure 1B). The first of two PCRs ( Figure 1A) creates a linear insert with plasmid sequences at both ends (see Supplementary Materials for methods and instructions for primer design). It is, however, relatively straightforward, efficient, and reliable. Overlap extension PCR cloning, described here, is not the first form of PCR-mediated cloning ( 8– 10). Recombinases are generally sold as proprietary components of cloning kits, so few consumers optimize the in vitro recombination reactions. TA cloning and LIC require end modifications that cannot be monitored by gel electrophoresis. The methods that are easiest to monitor and optimize ultimately prove the most reliable. The practical utility of any cloning method is predicated upon its reliability, rather than its convenience, price, or efficiency under optimum conditions. 
Numerous alternative approaches to PCR cloning ( 1) have been developed, including TA cloning ( 2), ligation independent cloning (LIC) ( 3– 4), recombinase-dependent cloning ( 5– 7), and PCR-mediated cloning ( 8– 10).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
